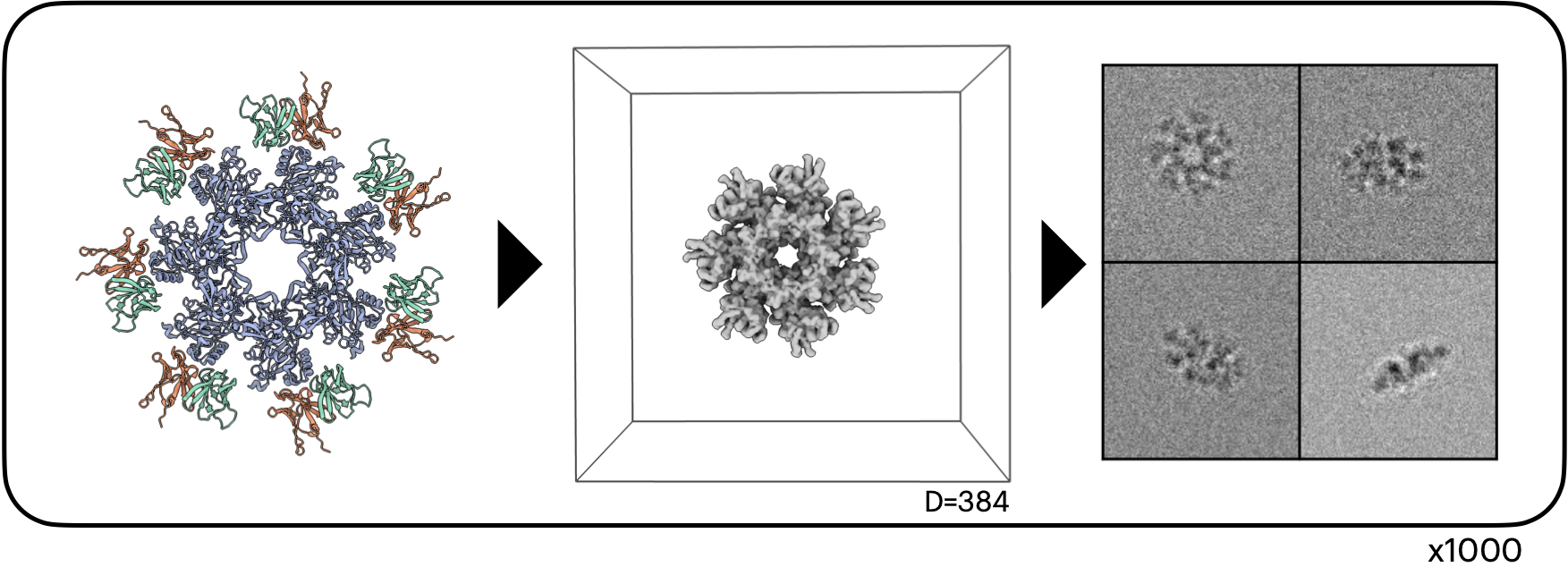
CryoHype successfully reconstructs the 1000 structures of Sim2Struct-1000
Abstract
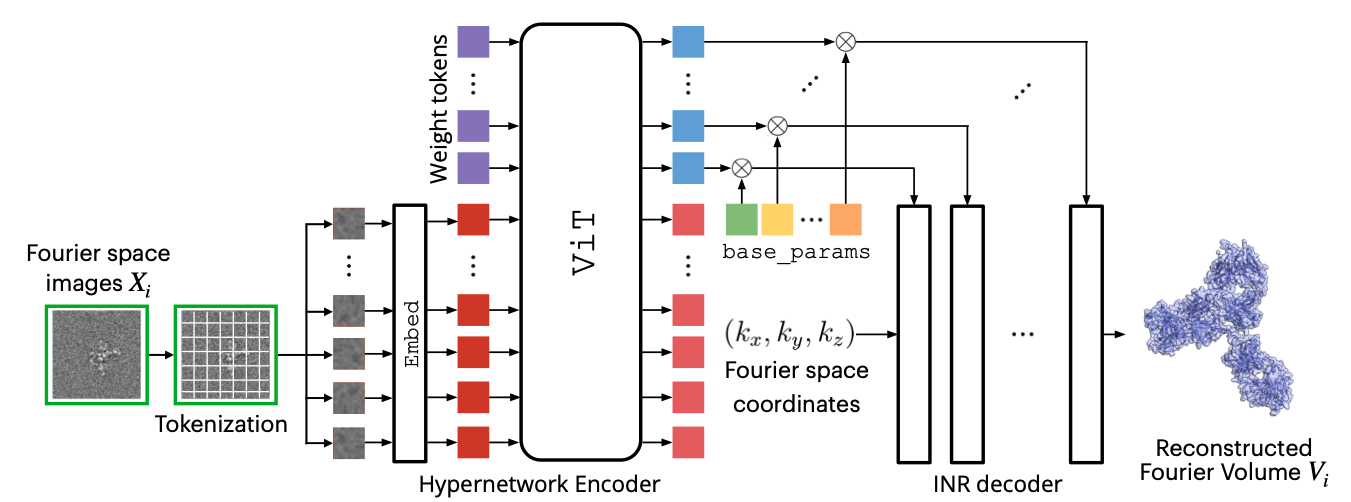
Cryo-electron microscopy (cryo-EM) is an indispensable technique for determining the 3D structures of dynamic biomolecular complexes. While typically applied to image a single molecular species, cryo-EM has the potential for structure determination of many targets simultaneously in a high-throughput fashion. However, existing methods typically focus on modeling conformational heterogeneity within a single or a few structures and are not designed to resolve compositional heterogeneity arising from mixtures of many distinct molecular species. To address this challenge, we propose CryoHype, a transformer-based hypernetwork for cryo-EM reconstruction that dynamically adjusts the weights of an implicit neural representation. Using CryoHype, we achieve state-of-the-art results on a challenging benchmark dataset containing 100 structures. We further demonstrate that CryoHype scales to the reconstruction of 1,000 distinct structures from unlabeled cryo-EM images in the fixed-pose setting.
BibTeX
@article{gu2026cryohype,
title={CryoHype: Reconstructing a thousand cryo-EM structures with transformer-based hypernetworks},
author={Gu, Jeffrey and Jeon, Minkyu and Ma, Ambri and Yeung-Levy, Serena and Zhong, Ellen D},
journal={Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition},
year={2026},
url={https://cryohype.cs.princeton.edu}
}